 Apply one method to properly segment the stack, e.g.Appreciate that this montage view is not suited for further analysis of the binary output.Observe that the different methods give different outputs.Appreciate that they yield different results. 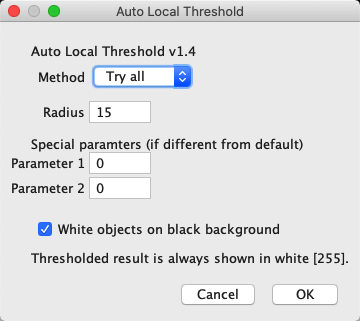
Apply one or more automated thresholding methods to this image.# Note that there exists a threshold_multiotsu function to handle cases with multi-peaks histograms add_image ( mean_thresholded2, name = 'mean_thresholded2', opacity = 0.4, colormap = 'magenta' ) # %% add_image ( mean_thresholded1, name = 'mean_thresholded1', opacity = 0.4, colormap = 'magenta' ) viewer. add_image ( manual_thresholded2, name = 'manual_thresholded2', opacity = 0.4, colormap = 'magenta' ) # Identify possible problems with this solutionįrom skimage.filters import threshold_mean thr1 = threshold_mean ( image1 ) print ( thr1 ) mean_thresholded1 = image1 > thr1 thr2 = threshold_mean ( image2 ) print ( thr2 ) mean_thresholded2 = image2 > thr2 # %% add_image ( manual_thresholded1, name = 'manual_thresholded1', opacity = 0.4, colormap = 'magenta' ) viewer. Thr1 = 25 thr2 = 75 manual_thresholded1 = image1 > thr1 manual_thresholded2 = image2 > thr2 viewer. max ())) print ( ' \n Min image2 %d max image2 %d' % ( image2. dtype ) print ( ' \n ', info_type ) print ( ' \n Min image1 %d max image1 %d' % ( image1. add_image ( image2, name = 'image2' ) # %% add_image ( image1, name = 'image1' ) viewer. # Instantiate the napari viewer and display the images Image1, axes1, scales1, units1 = open_ij_tiff ( '' ) image2, axes2, scales2, units2 = open_ij_tiff ( '' ) # %% Import napari import numpy as np from OpenIJTIFF import open_ij_tiff # %% In auto thresholding several methods produce acceptable results and one can get rid of selecting manual threshold values for each different image.Select window xy_8bit_nuclei_with_offset.tif.Select window xy_8bit_nuclei_without_offset.tif.Here, selecting Lower threshold level = 40 works.Selecting Lower threshold level = 20 now does not work (i.e. 
Selecting Lower threshold level = 20 is a good example.Open xy_8bit_nuclei_without_offset.tif. 
To summarize, in comparison to CellProfiler and ImageJ, quantification results in FoCo vary from the reference value less than 10 for both irradiated and control image sets indicating the reliability of quantifications.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
